Welcome Back!
Thanks for tuning in to the FAIR-CF newsletter - where the goal is to show how CF researchers of all stripes can make the most of public data.
Read the FAIR-CF manifesto here.
This month I’ve made some more changes to improve the reading experience.
The research summaries are different. The core elements remain (the goals, methods, results, and future directions are still discussed), but the style has changed. I want to tell the story of this research space, and show that under the surface, a global network of researchers are really working together on shared research projects - even if they have never seen or heard of each another before.
The summaries are news segments in three parts: a lead-in, the details, and the big picture. The hope is that somewhere along the way - maybe not in this edition, but the next one, or the one after that - these short research stories will inspire you to push forward with your own research in a new way. They may even help you make new personal connections. At the very least, they should stir your faith in the scientific enterprise, driven by an international community of diverse backgrounds, interests, and talents - yet ultimately, united in a common cause.
You can subscribe to FAIR-CF using the button below if you are not already. And make sure to share with your colleagues too!
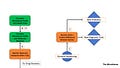
Research Map
As we develop a more detailed picture of the field, and discover new research projects, it becomes impossible to fit everything into one small square, so there are now multiple sections of the research map.
Map Key: Green = newly discovered research project, orange = previously discovered research project ; solid boxes involve research featured in this months study summaries. Blue circles are known data sources and blue diamonds are known translational endpoints. If you want to view the map in full screen, just click on the images.
News From the Field
Gut Biology
(A) miRNAs as Cancer Biomarkers
Human cells contain complex molecular networks of genes and proteins – and in cancer cells, these networks have gone awry. Scientists want to know why.
Researchers in Norway are focused on miRNAs, small compounds that regulate the translation of a cell’s genes into proteins. They found that certain miRNAs are more abundant in colorectal cancer tissue than surrounding cells. Several of these miRNAs are novel – they have not been previously studied in the cancer context and their function needs to be explored.
Featured Article: A comprehensive framework for analysis of microRNA sequencing data in metastatic colorectal cancer 🇳🇴

💾 Public Data: miRNA expression profiles (Databases: EGA, SRA, GEO)
With this information in hand, scientists may study the novel miRNAs to see how exactly they contribute to cancer, and also try disrupting these cancerous miRNAs as a potential form of therapy. In related news…
miRNAs can be biomarkers not just for colorectal cancer in general, but also for specific colorectal cancer complications. Chinese researchers have established that the miRNA ‘miR-31’ is more abundant in patients with lymph node metastasis than patients without. 🇨🇳
A similar study by a multinational team of researchers from Qatar, Egypt, and Saudi Arabia asked the same question about breast cancer: What miRNAs are associated with lymph node metastasis and how does their presence predict patient prognosis? 🇶🇦🇪🇬🇸🇦
Can miRNAs predict chemotherapy response too? Indeed, they can. Scientists in Seville, Spain have found that patients expressing higher levels of certain miRNAs responded better to chemotherapy. 🇪🇸
(B) The Gut Microbiome-Colorectal Cancer Connection
Now take a step back – what causes colorectal cancer in the first place? Scientists have reason to believe that the microbiome plays a role.
Scientists from China and the US are exploring the microbiome-cancer connection. Looking at the prior literature, the team identified a list of bacterial species that are more (or less) abundant in colorectal cancer samples than samples from regular patients. Looking at the protein-coding sequences of these bacterial species, the researchers came up with a list of nearly 100 bacterial proteins that could be contributing to cancer.
Featured Article: Prediction of Pathogenic Factors in Dysbiotic Gut Microbiomes of Colorectal Cancer Patients Using Reverse Microbiomics 🇨🇳🇺🇸

💾 Public Data: Protein-encoding sequences of enriched and depleted bacteria (Database: BioProject)
These are just predictions – they need to be tested in the lab – but any compound that is experimentally verified as a contributor to cancer may become a potential therapeutic target. And restoring a species that is lacking in the CRC microbiome could be a powerful probiotic strategy. In related news…
Another team of researchers in the States did a similar study, using public data on nearly 1000 tumor samples to identify a shared colorectal cancer ‘tumor microbiome’. They ultimately found 114 common tumor-associated microbes, and even built a web site that allows you to look at their results. 🇺🇸
What about other diseases? Scientists in China have shown that the gut microbiome of people with atherosclerosis (a type of cardiovascular disease) is altered too. 🇨🇳
Looping back to a familiar topic – miRNAs – American researchers have found that certain miRNAs are correlated with the relative abundance of bacteria (i.e., the % of the total microbiome that they make up). They too may play a role in shaping the tumor microbiome. 🇺🇸
C. Expanding the Probiotic Arsenal
Scientists want to identify new strains of probiotics and build the probiotic arsenal. The goal is to prescribe probiotic regimens that better suit individual patients.
Chinese Scientists compared the DNA of more than 100 separate strains of the probiotic bacterium Akkermansia muciniphilia. They were able to map out the genetic similarities and differences of the strains – and then arranged the strains on a family tree (phylogenetic tree), showing that they fell into three major lineages.
Featured Article: Comparative Genomics Revealed Wide Intra-Species Genetic Heterogeneity and Lineage-Specific Genes of Akkermansia muciniphila 🇨🇳

💾 Public Data: 112 A. muciniphilia genomes (Database: RefSeq)
With this information in hand, researchers may begin studying the functional differences between the strains more closely – and ultimately figure out which probiotic strains are best to treat specific diseases. In related news…
Another team of Chinese scientists recently conducted a similar study comparing A. muciniphilia genomes, but employed a different set of analysis techniques. 🇨🇳
Comparative genomics is not just useful for probiotics research. Scientists are studying the bacterium M. luteus, a common cause of hospital-acquired infections which may also have applications in the biotech industry. 🇨🇳
Taking a step back, before any ‘comparative genomics’ analysis can be done, bacterial strains need to have their genomes sequenced. Scientists in New Zealand sequenced Pseudobutyrivibrio xylanivorans MA3014, which helps cattle (and other ruminants) digest plants. 🇳🇿
D. Understanding the ‘Athlete’s Microbiome’
The ‘athlete’s microbiome’ is a known phenomenon – people who exercise regularly tend to show differences in their gut microbes.
To understand whether certain microbiome modifications might help athletes to train, perform, and recover, scientists at the University of Parma re-analyzed over 200 microbiome samples from a mixed population of athletes and non-athletes. Using a technique called shotgun metagenomics, they found three distinct microbiome profiles based on activity level (from non-athletic to highly athletic). They also showed that the athlete’s microbiome was richer in anti-inflammatory bacteria, and also home to more short fatty acid producing bacteria, which are thought to boost energy production.
Featured Article: Investigation of the Ecological Link between Recurrent Microbial Human Gut Communities and Physical Activity 🇮🇹

💾 Public Data: 207 Metagenomic samples from 6 published studies (Database: SRA)
While this study took a snapshot approach, looking at the athletic vs. the non-athletic microbiome, other studies may build on these findings by looking at how the microbiome changes over time in single individuals who adopt new habits. In related news….
What happens when sedentary individuals start to exercise – how does their microbiome change? For beginner athletes at least, the changes are pretty modest, a study conducted in Ireland and the UK has found (the authors suggest that major changes may only come with long-term activity). 🇮🇪🇬🇧
On a related note, another recent study monitored well-trained athletes taking part in a 5000km, 33 day rowing race across the ocean. Microbiome samples were collected before, during, and after the race – and in this case, more significant changes in the microbiome were observed. 🇮🇪
Scientists are studying the effects of diet on the microbiome as well – and not just for humans. An international research group studied that gut microbiome of 77 different mammalian species! 🇮🇹🇮🇪
Research Community
This month’s featured research and related studies involve researchers in 11 countries, including 5 US states
Spread the Word!
Thanks for reading! If you want to help us in our mission to help CF researchers make the most of public data, please share this newsletter with any colleagues who would be interested. Just press the button below to share…