Introduction
Thanks for tuning in to The Data Reuse Digest! In writing this newsletter, our goal is to uncover all the different ways that published scientific data can be used to drive research forward - with an eye on translational developments (new drugs, new clinical guidelines, new technologies).
We try to keep the writing plain and simple so that the newsletter can be useful to researchers of any field (not just bioinformatics) and the general public as well.
For researchers, especially younger researchers, this newsletter will show you what publishable work in the field looks like right now. It also hopes to encourage new kinds of research projects that push the field in new directions, towards new translational goals. For non-researchers, this newsletter will pull back the curtain to show what research actually looks like and why it matters.
If you are not a subscriber already, you can subscribe to the Data Reuse Digest here:
Bioinformatics Research Roadmap
Why does data reuse research matter? This road map shows how bioinformatics projects that gather together all kinds of published scientific data can advance research and produce important translational applications (new drugs, diagnostics, clinical guidelines and more)
Research News
(A) New Algorithms
Developing Computational Tools to See Biological Data in New Ways

The RNA molecule is fundamental to human life - orchestrating the activity of human cells and serving as the basis for therapies like the mRNA vaccines developed to halt the spread of COVID-19. This article discusses the design of RNA structure prediction algorithms - which predict the 3D molecular structure of an RNA molecule based on its sequence of nucleotides. These algorithms are similar to the popular AlphaFold algorithm that predicts the 3D structure of proteins from their underlying sequence of amino acids. AlphaFold is already being applied to design new drugs. Likewise, new and improved algorithms for RNA design will hopefully promote the development of more effective RNA-based therapeutics [Ward et al., 2023] 🇺🇸🇦🇺
Researchers have developed a new algorithm that takes an input DNA sequence and identifies interaction sites - places where transcription factors (TFs) that regulate gene expression are likely to bind. Algorithms that identify TF binding sites are useful for biomedical research, as mutations in these binding sites may play a role in various human diseases. The algorithm has been made accessible as a Python tool so that other researchers can use it in their own research [Mille et al., 2023] 🇫🇷
Another team constructed a machine learning model (using a random forest classifier) to identify the protein expression patterns that define different cell and tissue types in the human body - drawing on data from nearly 200 studies in the PRIDE proteomic database. The model is able to predict the cell or tissue type of a given sample based only on the proteins expressed in that sample. One possible application of this model is determining how well an organ engineered in the lab resembles a natural human organ of the same type. This makes it a valuable asset for regenerative medicine [Claeys et al., 2023] 🇧🇪
A core project of biological research is to figure out the function of every protein - and determine how protein sequence and structure relate to a protein’s functional abilities. A large international research team trained their sights on over 1000 different microorganisms, predicted the structure of 200,000 different microbial proteins, and mapped specific ‘protein folds’ (subcomponents of these proteins with defined 3D structure) to functions like ‘carbohydrate binding’ or ‘membrane transporter activity’. Interestingly, while some functions are accomplished by a very specific kind of protein fold (a one:one relationship), other functions can be accomplished by a variety of different protein folds (a one:many relationship). Another exciting aspect of this study is that the research team relied on a massive network of non-scientists to help them predict protein structures, partnering with the large citizen science initiative World Community Grid. [Leman et al., 2023] 🇺🇸🇵🇱🇳🇿🇫🇮🇩🇪
Often, one drug alone is not enough to kill cancer - but how do you know what combination of drugs will work best? Researchers have developed a new deep learning approach to predict synergistic drug combos for cancer treatment. The neural network classifier, named SYNDEEP by its designers, uses existing drug data (drug chemical structure, known human protein targets, etc.) as inputs to infer how multiple drugs will impact the body if administered at the same time [Torkamannia et al., 2023] 🇮🇷🇺🇸
(B) New Databases
Building databases to store and share biological knowledge

Mice are the most frequently used test animals in a variety of experiments. The data gathered from them has been distributed over numerous databases dealing with vastly different medicinal research topics. To provide easier access to this data, specifically for cancer research, the Mouse Tumor Biology database (MTB) was launched in 1998, containing data about pathobiology of cancer in genetically defined strains of mice. Through 2019-20, this database was improved in order to provide deeper insights into the genome of genetically engineered mice as well. It has recently been renamed as the Mouse Models of Human Cancer database (MMHCdb), and currently links up with many other databases to provide users with easy access to even more data [Begley et al., 2023] 🇺🇸
Extrachromosomal circular DNA (eccDNA), first discovered in the mid-1960s, are pieces of DNA found outside of chromosomes in the nuclei of eukaryotic cells. They are thought to interact with chromosomes and help regulate gene expression - but also tend to carry genes which are highly expressed in cancer cells, and may contribute to cancer progression. The eccDB database has been developed as a comprehensive repository that maps out eccDNA-chromatin interactions in cells across multiple species. It provides users with the ability to browse, search and analyze eccDNAs interactions or analyze unknown sequences for similarities with known eccDNA. [Yang et al., 2023] 🇨🇳
Continuing with the theme of regulating gene expression, cis-regulatory sequences, segments of DNA typically located upstream of a gene on the same DNA molecule, are well known for their regulatory role. Not all cis-regulatory elements in the genome are mapped - some are yet to be identified. A team of researchers has developed a new convolutional neural network (CNN) model which recognizes patterns in genomic sequences and predicts where the cis-regulatory sequences lie [Wei et al., 2023] 🇨🇳🇺🇸
Synthetic Biology involves re-engineering organisms to give them new abilities - for example, modifying bacteria so that they can clean pollutants from drinking sources. Synthetic biology experiments employ various data processing, computational modeling and artificial intelligence tools. Several repositories of analytical tools have been created but none of them encompass all the tools available to this date. To address this issue, researchers have created SynBioTools, curating all available databases and related data from published reviews to aid fellow synthetic biologists [Cai et al., 2023] 🇨🇳🇨🇭
Transposable Elements (TEs), otherwise known as ‘jumping genes’, are DNA sequences that are able to move around to locations in the genome. Their movement can impact the expression of other genes and in some cases may contribute to diseases like cancer. Analyzing the genome for the presence and movement of TEs can provide new insight into cellular function and disease risk. TrEMOLO, short for Transposable Element MOnitoring with LOng-reads, is a new software that takes advantage of long-read sequencing data to identify TEs and their abundance in the human genome [Mohamed et al., 2023] 🇫🇷
(C) Bioinformatics Analysis
Bioinformatics tools applied to answer key biological questions

Many years of research have gone into the study of asthma, but there are still new ways to investigate the disease. This recent study aimed to establish the key differentially expressed genes (DEGs) that distinguished the airway cells of asthma patients from those of healthy individuals. The authors used a technique called WGCNA to identify modules of correlated genes that are active in asthma cells, and pinpoint the ‘hub genes’ that drive the expression of these modules. Through this analysis, the researchers identified CYCS as a gene that could help predict asthma prognosis [Li et al., 2023] 🇨🇳
Continuing with the theme of chronic respiratory disorders, a newly published study aimed to support the diagnosis and treatment of lung cancer - specifically non-small cell lung cancer, which accounts for 80-85% of total lung cancer cases. Using gene cohorts for lung adenocarcinoma (LUDA) and lung squamous cell carcinoma (LUSC) obtained from The Cancer Genome Atlas (TCGA), researchers performed bioinformatic analysis to identify the gene ZWINT as one that likely contributes to non-small cell lung cancer [Cao et al., 2023] 🇨🇳
The COVID-19 pandemic, though under good control, still poses threats to public health. There are unanswered questions about its tendency to exacerbate other illnesses and to cause ‘long-COVID’ symptoms. A recent study sheds light on the relatedness of COVID-19 with Intracranial Aneurysm (IA) on a molecular level. Researchers identified differentially expressed genes that are implicated in COVID-19 and IA. The researchers also drew on information from public databases to identify drugs that may target the protein products of these differentially expressed genes and potentially be used to treat both diseases [Snigdha et al., 2023] 🇧🇩
Another study focuses on the pathobiology of Parkinson's Disease (PD). Researchers compared gene expression in substantia nigra tissue (a region of the brain implicated in Parkinson’s disease) to gene expression in the blood of Parkinson’s patients, drawing on multiple datasets from the Gene Expression Omnibus (GEO). They identified several differentially expressed genes in the substantia nigra tissue and blood samples, and like the COVID/IA study, identified drug compounds that could potentially target the protein products of these genes [Elango et al., 2023] 🇸🇦🇦🇪
(D) Commercial News
Bioinformatics-based businesses driving new drug development

Absci Corporation and Aster Insights (formerly M2GEN), two companies in the AI and bioinformatics space, have partnered up to design new cancer drugs. Aster is bringing the biological data, sharing its massive AVATAR database of genetic and clinical information from over 350,000 cancer patients. Absci will search through this database, using its own AI drug creation platform, to discover potential targets for new immunotherapy drugs 🇺🇸🇨🇭
Aster Insights also announced results of another partnership with California-based biotech Twist Bioscience. Twist has developed a new screening panel - which uses whole exome sequencing to detect hundreds of cancer-associated genes that are not typically captured by other screening platforms. The panel is especially well suited to identify cancers that are driven by copy number variation - a situation where a single gene can be copied multiple times in the genome, amplifying its biological impact. The new technology, labeled the ‘AsterExome’ panel, will be integrated into Aster’s AVATAR program 🇺🇸
Outside of cancer drug development, Israeli AI firm Identifai-Genetics is developing a new platform for early, non-invasive detection of genetic disorders. The company closed 3.3 million dollars in funding to support its efforts. The Identifai platform can detect genetic disease in a developing fetus based only on a blood test from the mother. According to the company, existing non-invasive screening methods detect less than 10% of known genetic disorders. The Identifai platform is designed to be much more comprehensive 🇮🇱
Often, firms focused on data science serve in a consulting capacity for other companies. This is the case for Excelra, which partners with leading pharmaceutical and biotech companies to help them use biological data to make new drug discoveries. Excelra recently acquired the consulting firm BISC Global, which will expand the strength of Excelra’s services in the areas of bioinformatics, biostatistics, AI, and machine learning. 🇮🇳🇧🇪
In the realm of new technology, the global market for RNA sequencing and analysis is expanding, with an expected growth of 14% in the next decade. New technologies in development include a platform that is capable of capturing both mRNAs and miRNAs in single cells at the same time. Technologies like this one that simultaneously capture different kinds of genetic information will provide a more complete understanding of the genetic factors that drive disease and contribute to differences in patient outcomes 🌎
Research Community
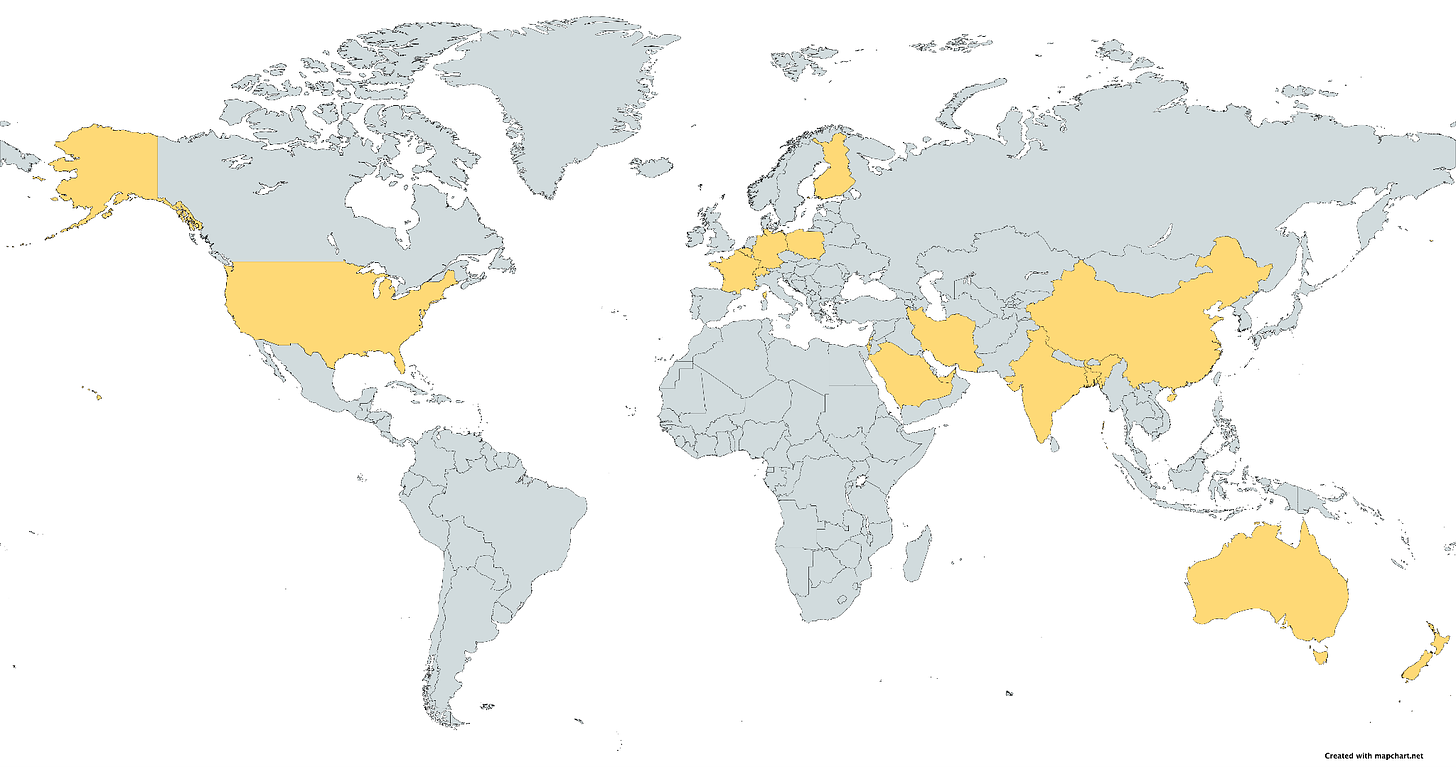
This month’s new studies involved 16 countries around the world
Spread the Word!
Thanks for reading! If you want to help us in our mission to show how researchers can make the most of public data, please share this newsletter with any colleagues who would be interested. Just press the button below to forward the newsletter along