Introduction
Thanks for tuning in to The Data Reuse Digest! In writing this newsletter, the goal is to uncover all the different ways that scientific data can be used to drive research forward - ultimately with an eye on translational developments (new drugs, new clinical guidelines, new technologies, etc.).
For researchers, especially younger researchers, the idea of this newsletter is to show what successful, publishable work in this area of analyzing big biological datasets looks like right now. At the same time, it also aims to encourage new kinds of research projects that push the field in new directions, towards new translational goals. For non-researchers, the idea is to pull back the curtain to show what research work actually looks like, and why it matters for society at large.
You can subscribe to the Data Reuse Digest here if you are not a subscriber already:
SPECIAL NOTES:
This newsletter has become a team effort! Sharon Tribhuvan, a friend and fellow science writer who studies biochemistry at the University of Delhi, contributed to the research summaries this month
In news related to this newsletter, I wanted to advertise an upcoming summer bioinformatics course, organized by the Stanton Lab at Dartmouth College and hosted at the Mount Desert Island Biological Labs in beautiful Bar Harbor Maine. The two-week intensive course is all about training researchers to get comfortable using public data and applying bioinformatics techniques in their everyday research. In the past, we have had graduate students participate from all across the US and even internationally. Scholarship support is available. Please consider the opportunity and pass along to other researchers in your orbit - you can share this link: https://mdibl.org/course/reproducible-and-fair-bioinformatics-analysis-of-omics-data-2023/
Research Maps
The research maps showcase the central projects that researchers are carrying out in the field. All of the featured studies below are represented by the letters on the map.
New Project: Gather large volumes of genomic data from the population to uncover disease-causing genetic variants so that their biology may be explored further and drug candidates developed or repurposed for disease treatment.
Established Projects from past editions:
Research News
Large International Population Study Identifies New Drug Targets for Stroke
🏘Reader Keywords: #Stroke #New Drug Targets #Drug Repurposing #Genome-Wide Association Study #Schizophrenia
Featured Article: Stroke genetics informs drug discovery and risk prediction across ancestries 🌍
Combining genetic data from many patients - across the world - can help scientists discover new ways to treat medical conditions. Take stroke - for example - which is the second leading cause of death worldwide. An international team of scientists have gathered data from 110,000+ patients and analyzed that data to identify new drug targets and opportunities for drug-repurposing. This study is extra special because it included a large portion of non-European participants. Previous studies like this one had mainly focused on people of European ancestry

💾 Data Source: Genetic data combined from multiple patient cohorts
This work is an example of a genome-wide association study (GWAS), where the genetic profile of people who have had a stroke and people who have not is compared to see how they differ. In addition to genomic data, the researchers also had transcriptomic data on hand for many of the participants. So not only could they see what genetic variants are associated with stroke - they could also see whether genes are contributing to stroke by being relatively over-expressed or under-expressed. Lower expression of the ICA1L gene in a region of the brain called the DLPFC, for example, was found to increase stroke risk.
Not stopping there, the researchers ran a 'genomics-driven drug discovery' protocol - an analysis approach that has become popular in the last few years. It works, in part, by taking the stroke-associated genes identified in the study and scanning public databases (such as ‘DrugBank’) to ID existing drugs known to target those same genes. In the end, the research team put their finger on 6 genes that they believe could be drug targets to prevent or alleviate stroke symptoms (F11, KLKB1, PROC, GP1BA, LAMC2 and VCAM1). Two of these genes (F11 and PROC) are targeted by drugs that are already under development to treat or prevent stroke.
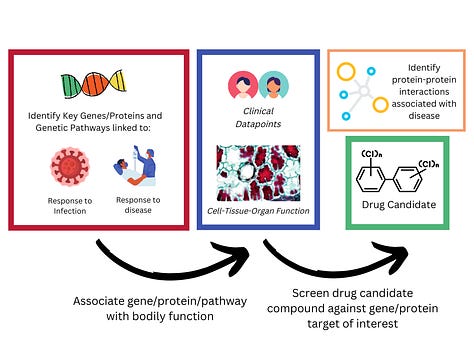
Other scientists are finding genetic variants that define different diseases (1) and testing the biological effects of these variants in animal models (3) - or looking at how expression of certain proteins in various tissues is associated with disease risk (2)
🗺Translational Path:
Similar Article: Another related study examined schizophrenia in an East Asian population (a previously understudied group), looking for genetic variants that distinguished people with schizophrenia from those without. 🌍
Cited (1): Building on knowledge of the genetic variants associated with schizophrenia, scientists have looked at the levels of certain proteins expressed in the brain and spinal fluid - expression of certain proteins (C4A/C4B, ACP5, CNTN2, PLA2G7) was associated with a lower risk of schizophrenia, and others (TIE1, BCL6, MICB) with an increased risk. 🇨🇳
Cited (1): Focusing in on a specific SLC39A8 gene variant associated with schizophrenia, researchers built a mouse model with the variant to assess its biological effects. They identified several traits that defined mice with this variant (compared to control mice without the variant): less zinc in the brain and a lower dendritic spine density - which may play a role in schizophrenia symptoms. 🇨🇳
Researchers Scan South Asian Population for Disease and Drug Resistance Variants
🏘Reader Keywords: #South Asia #Genotyping #Disease-Causing Genetic Variants #Pharmacogenomics #Ethnic Variation #GWAS #Multi-Omics #DNA Methylation #Transcriptomics #Biological Database
Featured Article: Analysis of clinically relevant variants from ancestrally diverse Asian genomes 🇸🇬
The human genome project. The landmark effort to sequence the human genome has told us volumes about ourselves: revealing secrets about human health and identifying new opportunities to treat disease. But early efforts to report genomic information have not covered the whole global population equally. Thankfully, there are projects underway to address these past limitations and capture the genomic variation of all humankind.

💾 Data Source: Individuals participating in six different studies provided genetic data
Here is one - a large team of researchers in Singapore have analyzed nearly 10,000 genomes from the people of South and East Asia, a previously under-represented group. The researchers were able to identify a number of disease-causing genetic variants that are common in this population, as well as a number of variants that are likely to interfere with the activity of drugs. These variants, and their known implications, had been discovered in past genetic studies - so this work is the culmination of many different papers.
Other scientists are comparing the genomes of different ethnic groups in South Asian populations (1), performing genome-wide association studies to find genetic variants associated with disease (2), and building interactive databases to capture genetic profiles and make them accessible to study by the research community (3).
🗺Translational Path:
Similar Article: This group compared the genomes of nearly 5000 people from three different ethnic groups (Chinese, Malays, Indians) in Singapore, identifying numerous novel genetic variants. 🇨🇳🇸🇬🇺🇸🇦🇺🇳🇿
Cited (1): Researchers performed a genome-wide association study to identify variants associated with Type II diabetes in a South Asian population. They identified 21 novel genetic variants associated with the disease. 🌍
Cited (1): Another research group compiled ‘multi-omics’ data on Asian pregnant woman - combining three forms of data (genotyping, DNA methylation, and transcriptome profiling) together in a single database. The study authors were able to demonstrate genetic variation between different ethnic groups. 🇸🇬🇨🇦🇦🇺🇫🇮🇳🇿
Scientists Establish Causal Links Between Pollutants and Respiratory Disease
🏘Reader Keywords: #Pollutant Exposure #Lung Disease #GWAS #Lung Cancer #Lipid Metabolism #D-Limonene #Asbestos #Silica
Featured Article: Consequences of exposure to pollutants on respiratory health: From genetic correlations to causal relationships 🇮🇹
Pollution has a long history of association with disease, and this history comes with potent images: acrid smoke belching from the mills of an industrializing world, or a hazy smog of automobile exhaust blotting out the sun on the city skyline. Lung disease is indeed one of the top 5 global causes of death. But by what mechanism do these pollutants of the air damage the lungs? What genetic variants determine why people respond more severely to pollutants than others?

💾 Data Source: Genetic data gathered from the UK Biobank
Researchers have found a clever way to answer this question with public data. In this study, researchers drew on 170 GWAS datasets, which link genetic variants to individual traits. Most GWAS studies ask the same question: is there a certain genetic variant associated with disease? But this study went a step further and made predictions about how pollutants are interacting with certain genetic variants to cause disease.
Take, for example, the question of whether being in a workplace with lots of diesel fuel exhaust contributes to respiratory disease - and what genetic variant is responsible for this effect. First, the researchers noted that people who work with diesel exhaust are more likely to have respiratory diseases. Then, they looked at genetic variants. If a genetic variant is causing respiratory disease in people who work in these conditions, then you should see it often in those individuals who work in these conditions and do develop respiratory disease, but not often in people who work in these conditions but don’t develop respiratory disease. The researchers were able to establish causal relationships between a number of pollutants and respiratory illness in this way.
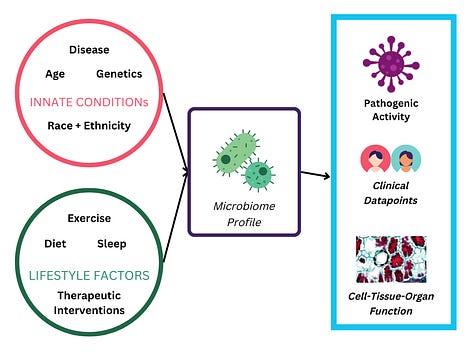
Other researchers are studying the causal link between air pollutants, carcinogens, and cancer risk (1,3). Scientists are further investigating the biological mechanisms by which pollutants cause disease, and even considering therapeutic interventions to alleviate pollutants’ negative effects (2).
🗺Translational Path:
Similar Article: Researchers assessed the impact of exposure to particulate matter in the air on lung cancer risk. They found that long-term exposure to air pollution may increase the risk of cancer. 🇨🇳
Cited (1): Researchers find that exposure to particulate matter in the air causes alterations in lipid metabolism. Furthermore, ingestion of D-Limonene was shown to alleviate these changes in lipid metabolism, and may be a useful preventative compound to ward off lung cancer if it proves successful in further trials. 🇨🇳🇺🇸
Cited (1): Researchers in China wanted to estimate trends in exposure to carcinogens in the workplace from the 1990s to today. Asbestos and silica were the most common carcinogens that workers were exposed to - and the researchers found that the incidence of lung cancer that can be attributed to occupational carcinogens increased significantly from 1990 to 2019. 🇺🇸
Researchers ID Genetic Variants That Impact Expression of Alzheimer’s Disease-Related Proteins
🏘Reader Keywords: #Alzheimer’s Disease #GWAS #Biomarker Discovery #Amyloid β 42 #Phosphorylated Tau #miRNAs #Gene Expression
Featured Article: Genome-wide meta-analysis for Alzheimer's disease cerebrospinal fluid biomarkers 🌍
Alzheimer’s disease is one of the most well-known and extensively studied neurodegenerative diseases, yet medical treatment is still elusive. Researchers have, thus far, elucidated several risk factors that can predict whether one develops the disease later in life. Perhaps these studies also may provide hints at potential treatments?
💾 Data Source: Combined genetic data from 16 European cohorts
An international group of researchers studied the genome to identify specific biomarkers that could aid in early detection and possibly even in the treatment of Alzheimer’s disease.
Amyloid β 42 (Aβ42) and phosphorylated Tau (pTau) are two proteins specific to AD that are found in the cerebrospinal fluid (CSF) of affected individuals. The study, comprising over 13000 patients in various stages of disease progression, focused on identifying and characterizing novel genes/loci that affect the levels of Aβ42 and pTau in patients.
Researchers were able to identify 2 loci (CR1 and APOE) that affect expression of Aβ42 and 4 loci (BIN1,GMNC, C16orf95 and APOE) that affect expression of pTau, 3 of which (CR1, BIN1, and C16orf95) are novel. The researchers also divided patients into clusters based on the level of expression for both Aβ42 and pTau, and suggest that divergent routes of treatment could be beneficial for these different clusters for patients. For example, individuals in one cluster would benefit more from treatments targeted at amyloid formation, while those in a second cluster would benefit more from treatments targeted at the immune system.
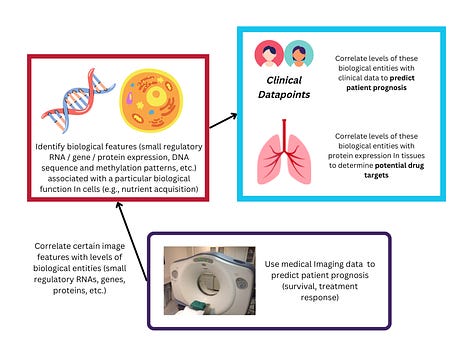
Other researchers are performing similar genome-wide association studies for Alzheimer’s (1) - and exploring the biology of Alzheimer’s further by identifying the patterns of gene (3) and miRNA (2) expression that define the disease.
🗺Translational Path:
Similar Article: Another group of researchers performed a similar genome-wide association study to find genetic variants that influence Aβ42, pTau, and Tau levels in the cerebrospinal fluid. 🇺🇸
Cited (1): Scientists identified miRNAs - small gene-regulating molecules - in the body fluids of Alzheimer’s disease patients and proposed several of these miRNAs as potential biomarkers for the disease because of their tendency to interact with Alzheimer’s related genes. 🇸🇮
Cited (1): At the gene level, researchers have found that two genes - ALCAM and BBX - were differentially expressed in Alzheimer’s patients compared to people without the disease. 🇺🇸
Research Community
This month’s featured research involved researchers in more than 9 countries (including several large international teams)
Spread the Word!
Thanks for reading! If you want to help us in our mission to show how researchers can make the most of public data, please share this newsletter with any colleagues who would be interested. Just press the button below to forward the newsletter along