Introduction
Thanks for tuning in to The Data Reuse Digest! In writing this newsletter, our goal is to uncover all the different ways that published scientific data can be used to drive research forward - with an eye on translational developments (new drugs, new clinical guidelines, new technologies).
We try to keep the writing plain and simple so that the newsletter can be useful to researchers of any field (not just bioinformatics) and the general public as well.
For researchers, especially younger researchers, this newsletter will show you what publishable work in the field looks like right now. It also hopes to encourage new kinds of research projects that push the field in new directions, towards new translational goals. For non-researchers, this newsletter will pull back the curtain to show what research actually looks like and why it matters.
If you are not a subscriber already, you can subscribe to the Data Reuse Digest here:
Bioinformatics Research Roadmap
Why does data reuse research matter? This road map shows how bioinformatics projects that gather together all kinds of published scientific data can advance research and produce important translational applications (new drugs, diagnostics, clinical guidelines and more)
Research News
(A) New Algorithms
Developing Computational Tools to See Biological Data in New Ways

Many cancer drugs are designed to target proteins called kinases. Sometimes, however, drugs designed to interact with a specific kinase will hit other unintended kinase proteins, causing dangerous side effects. Computational biologists at the National Cancer Institute have developed an algorithm that can identify subtle variations that distinguish one kinase from the rest - enabling the development of more selective anti-cancer drugs [Zhang et al., 2023] 🇺🇸🇮🇱
Scientists in Germany have developed an AI model called SCEMILA that can identify different kinds of acute myeloid leukemia tumor cells in images of human blood. The goal is to put this algorithm in the hands of clinicians and assist cancer diagnosis. Notably, the researchers describe their model as an ‘Explainable AI’ tool - one where the decision making process of the AI model is transparent [Hehr et al., 2023] 🇩🇪
Analyzing gene expression is a very common way to understand the activity of cells. For example, researchers can expose human cells to drug treatment and see how this changes gene expression. A new machine learning algorithm developed by researchers in Qatar and the United States allows researchers to view gene expression in a new way. With the algorithm, it is possible to see not only what genes are expressed, but also where in the cell they are being expressed [Musleh et al., 2023] 🇶🇦🇺🇸
Human organs are made up of large, diverse populations of cells, and understanding how organs work (or don’t work, in the case of disease) ultimately requires studying the interactions of individual cells. A new computational method developed by researchers in Germany employs graph neural networks to infer cell-cell interactions from single-cell sequencing and cell imaging data [Fischer et al., 2023] 🇩🇪
A common technique to measure a cell’s protein production is mass spectrometry (MS). MS involves striking a sample of proteins with a high energy electron beam - causing the proteins to break up into electrically charged fragments. Like a puzzle, researchers can piece together what proteins are present based on the fragments generated by MS.. Unfortunately, this puzzle is not always so easy to put together. A fragment could map to multiple different proteins, making researchers unsure which protein is actually present in the sample. To address this issue, Researchers from Singapore have developed a new tool called ProInfer. ProInfer makes use of prior knowledge about protein complexes (groups of proteins that are known to interact) - if one of the fragments detected by MS is part of a protein complex, and other proteins from that complex were detected in the sample, then it is likely that the fragment belongs to the complex protein rather than its other possible matches [Peng et al., 2023] 🇸🇬
(B) New Databases
Building databases to store and share biological knowledge

The behavior of a cell is determined by a series of complex underlying interactions between each gene and the ‘transcription factors’ that influence its expression. The study of these interactions, collectively known as Gene Regulatory Networks (GRNs) can provide a better understanding of the gene expression patterns associated with disease. Network Zoo (or netZoo for short) is a set of new software methods designed to help researchers infer new GRNs from multi-omics data (genomic, proteomic, metabolomic, and more) and analyze their activity [Guebila et al., 2023] 🇺🇸🇳🇴🇭🇰🇹🇼🇳🇱
Popularly known as ‘junk DNA’ until recently, non-coded ribonucleic acids (ncRNAs) have recently been shown to play critical roles in regulating gene expression. Long ncRNAs (having >200 nucleotides) are key factors in various diseases and play an important role in cell differentiation and development. lncHUB2 is a recently launched, full-stack, web-based application as well as an Appyter. It produces reports about various human lncRNAs by gathering data from other public databases and web-applications. It predicts lncRNA functions as well based on gene-gene co-expression correlation data [Marino et al., 2023] 🇺🇸
Centromeres, a component of DNA that participates in cell division, is essential for the transfer of genetic information through generations. The analysis of centromere architecture can provide useful insights about genome stability, cell division, and disease development. Centromeres exhibit extra-long tandem repeat (TR) units, referred to as monomers, which are further organized into higher-order repeats (HORs). Analysis of HORs, known as Centromere Annotation, gives insights about the structure and evolution of centromeres within and between species. HiCAT, short for Hierarchical Centromere structure AnnoTation, generates blocks, graphs, and other data points detailing the properties of centromeres [Gao et al., 2023] 🇨🇳🇳🇱
Traditional Chinese Medicine extensively utilizes plants, herbs and spices for the treatment of various diseases. A mechanistic understanding of the treatment, however, remains elusive due to lack of understanding of the exact chemical compounds involved in treatment and the underlying molecular mechanisms that they involve. IGTCM is an integrative genome database containing all the genetic information available, so far, on the plants and herbs used in TCM. This enables a better understanding of TCM for further drug discovery and also genetic improvement of TCM plants (some of which have assumed endangered status) through molecular breeding [Ye et al., 2023] 🇨🇳
Unconventional T cell receptors (TCRs) are biologically unique - they go against the well-established TCR-peptide-MHC paradigm and employ other MHC class Ib or MHC-like molecules like lipid moieties, etc., in their immune response. The fine-grained recognition pattern of antigens by unconventional TCRs is important for early life development and activation. Due to their small size and the variety in unconventional TCR sequences, their analysis and study has been tricky so far. For this purpose, UcTCRdb has been published as a comprehensive database that contains all published unconventional TCR sequences. It enables users to browse, download and analyze sequences and perform other operations as needed [Dou et al., 2023] 🇨🇳
(C) Beyond the Bench
Using data gathered outside the laboratory to help diagnose, treat and prevent disease

Convolutional Neural Networks (CNNs) have been used extensively to map the progression of dermatological and pathological diseases by analyzing spatial data such as images. Due to its strong image recognition capabilities, researchers have constructed a prediction model to accurately predict the chances of psoriatic arthritis (PsA) in patients already diagnosed with psoriasis (PsO), to provide them early examination and treatment that can prevent irreversible disease progression [Lee et al., 2023] 🇹🇼
Researchers have argued for the broader use of ‘digital phenotyping’ technologies in clinical trials, relying more on wearable devices like smart watches to track patient data. Up to now, researchers have largely relied on smartphones due to concern about the data quality from other personal devices. But as this review notes, the quality of data gathered by wearable devices has improved significantly, and recent clinical trials are starting to incorporate them [De Boer et al., 2023] 🇺🇸
Speaking of wearable devices, a recent study in the US used wearables to track the burden of influenza-like illness in the population. The participants reported any flu-like symptoms and submitted data including their total daily steps, active minutes, sleep quality, and resting heart rate. The participants who did report influenza-like symptoms were distinguished by their significantly reduced steps and active minutes, as well as increased sleep and higher resting heart rate. These results support the use of wearable data to help at-risk individuals seek medical care early if they come down with the flu - or even to track (and limit) the spread of an infectious disease in the population [Hunter et al., 2023] 🇺🇸🇨🇭🇬🇧
Researchers are moving beyond individual determinants of health to explore the impacts of social networks. Social connectedness is well known to reduce mortality in a wide range of conditions. Drawing on Facebook connection data from all 3142 counties in the mainland United States, this study found that as social connectedness rises, the prevalence of depression falls [Beauchamp et al., 2023] 🇺🇸
Another social media-based study tracked the mental health of social media users in Japan during the COVID-19 pandemic. The researchers used a machine learning algorithm called latent semantic scaling (LSS) to detect when levels of emotional distress are increasing in the population and what kinds of individuals are at highest risk. The algorithm could be used to identify people who are in danger of mental health crises and allow mental health professionals to intervene early [Ueda et al., 2023] 🇺🇸🇯🇵
(D) Bioinformatics Analysis
Bioinformatics tools applied to answer key biological questions

Machine learning methods can be used to predict remission in cancer patients - and identify the genetic features that predispose someone to better or worse clinical symptoms. In this study, researchers found a suite of genetic variants that predict survival in leukemia, drawing on data from a cohort of 1000+ patients [Eckardt et al., 2023] 🇩🇪
To treat all cancer patients successfully requires understanding how factors like age influence tumor behavior. A research team has examined over 10,000 tumor samples in adults and children, developed a neural network classifier to distinguish different cancer subtypes, and identified broad features that distinguish adult from childhood cancer [Comitani et al., 2023] 🇨🇦🇦🇺🇬🇧
Another key objective of cancer research is to distinguish invasive from noninvasive tumors - so that the former may be diagnosed and treated more aggressively. Researchers have run a quantitative proteomics experiment - comparing protein levels in invasive pituitary cancer to noninvasive pituitary cancer. They found that a certain protein, SLC2A1, was very well correlated with invasiveness and may serve as a biomarker for invasive pituitary cancer [Zhang et al., 2023] 🇨🇳
Grouping patients into distinct subgroups is useful for other conditions too. Researchers analyzed metabolite production in a cohort of obese individuals and used a neural network approach called self-organizing maps to define five distinct groups of patients (‘metabotypes’) based on their metabolite production. Certain metabotypes were found to be less responsive to bariatric surgery, indicating that additional or alternative treatments may be useful [Lappa et al., 2023] 🇸🇪🇳🇱🇩🇰
In a related effort, a large international team of researchers used a deep learning approach (Multi-omics variational autoencoders) to identify associations between the characteristics of individuals with type II diabetes and their response to different drugs. Patient characteristics included genetic variants, gene and protein expression, metabolite production, microbiome composition, diet, and a wide variety of clinical features (blood work, medical history, etc.) [Allesøe et al., 2023] 🌍
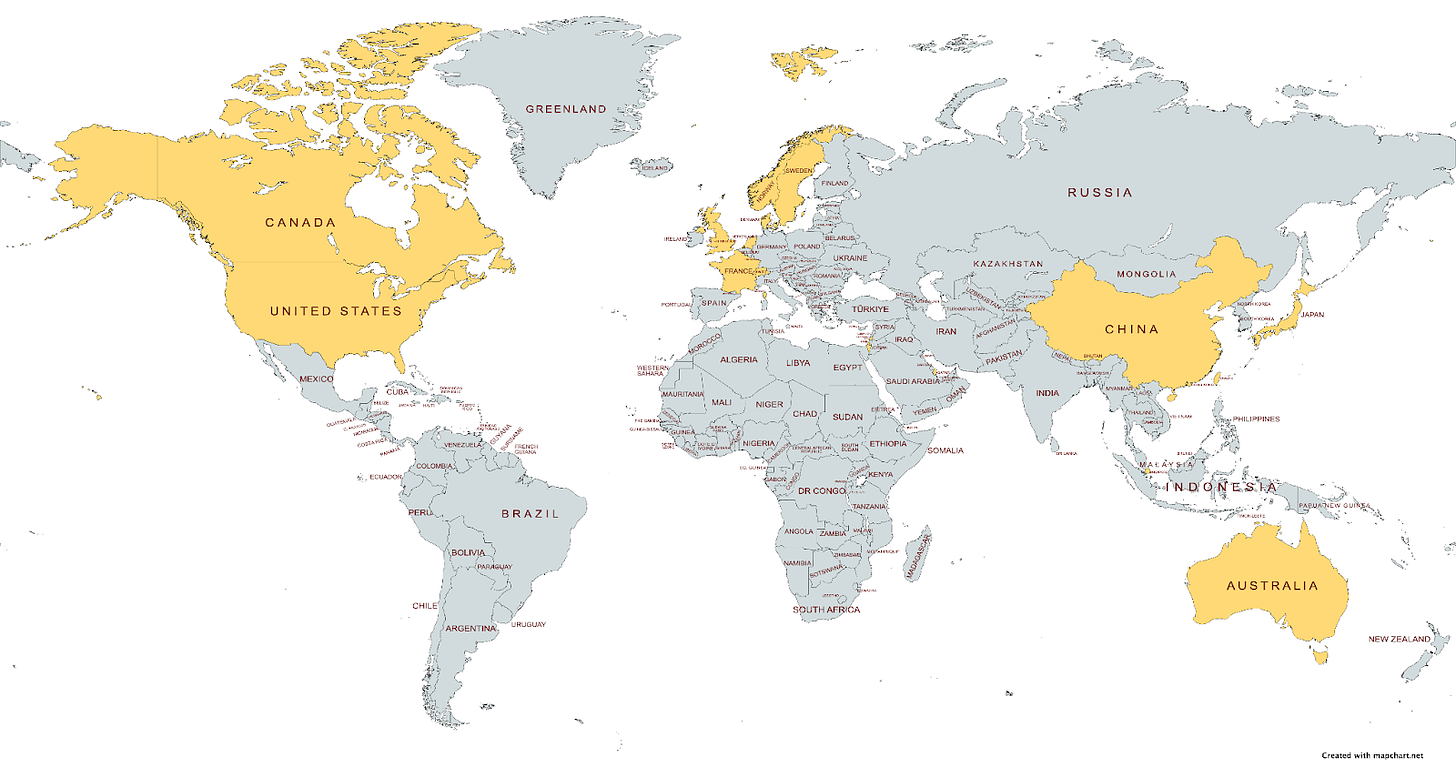
Research Community
This month’s new studies involved 17 countries around the world
Spread the Word!
Thanks for reading! If you want to help us in our mission to show how researchers can make the most of public data, please share this newsletter with any colleagues who would be interested. Just press the button below to forward the newsletter along