Introduction
Thanks for tuning in to The Data Reuse Digest! In writing this newsletter, the goal is to uncover all the different ways that published scientific data can be used to drive research forward - ultimately with an eye on translational developments (new drugs, new clinical guidelines, new technologies, etc.).
I try to keep the writing plain and simple so that the newsletter can be useful to researchers of any field (not just bioinformatics) and members of the general public as well.
For researchers, especially younger researchers, the idea of this newsletter is to show what successful, publishable work in the field looks like right now. At the same time, it also aims to encourage new kinds of research projects that push the field in new directions, towards new translational goals. For non-researchers, the idea is to pull back the curtain to show what research work actually looks like, and why it matters for society at large.
If you haven’t already, you can subscribe to the Data Reuse Digest here:
Research Map
The research map gives a brief roadmap to research in the field. All of the featured studies below are represented the letters on the map.
Research Project: Use published data to identify genes and proteins that are associated with disease and screen drug candidates to target them.
Research Project: Use published data to determine all the factors - positive or negative - that influence the microbiome and impact bodily function
Want to learn more about microbiome research? You can subscribe to our companion newsletter Mastering the Microbiome - just follow this link.
News from the Field
(A) Scientists Find Flu-Activated Genetic Pathways
🏘Reader Keywords: #Influenza A #Transcriptomics #WGCNA #Hub Genes #Inflammation #Gastric Cancer #Lung Cancer
Featured Article: Identification of Critical Genes and Pathways for Influenza A Virus Infections via Bioinformatics Analysis 🇨🇳
How does the body respond to the flu? Scientists are working to identify the genetic pathways in human cells that are activated by the influenza A virus (a common flu pathogen).

💾 Public Data: Cancer Gene Expression Datasets (GEO)
Researchers gathered data from 8 public gene expression studies, identifying which genes were turned on or off after infection with Influenza A virus (which is a common cause of the flu). They used a popular bioinformatics technique called Weighted Gene Co-Expression Network Analysis (WGCNA) to see which genes were turned on or off together across experimental samples. These ‘modules’ of correlated genes may work together and contribute to some part of the biological response to infection. After identifying these gene modules, the researchers went on to determine which modules were most active at different time points of infection - and what genes in the modules were likely to be driving their activity (‘hub genes’). They came up with a list of hub genes, including NDRG4, which is known to play a role in inflammation.
Building on these results, further studies may explore whether developing drugs to target genes like NDRG4 would help alleviate severe symptoms of the flu and other viral illnesses.
🗺 Translational Path:
Similar Article: Identify gene modules and hub genes (COL1A2, COL1A1, COL4A1, THBS2, ITGA5) associated with gastric cancer [Cao et al., 2018] 🇨🇳
Cited By: Verify experimentally that THBS2 gene expression is significantly higher in gastric cancer tissue and that elevated THBS2 expression is associated with poor prognosis [Zhang et al., 2022] 🇨🇳
Cited By: Show that elevated PSMD2 expression is associated with poor prognosis and infiltration of immune cells in lung cancer [Zhao & Lu, 2022] 🇨🇳
(B) The Shared Biology of COVID-19 and Rheumatoid Arthritis
🏘Reader Keywords: #Rheumatoid Arthritis #COVID-19 #Transcriptomics #Hub Genes #Drug repurposing #IPF #COPD #Diabetic Cardiomyopathy #Terpenoids
Featured Article: Bioinformatics and System Biology Approach to Identify the Influences of COVID-19 on Rheumatoid Arthritis 🇨🇳🇮🇳
Why is rheumatoid arthritis (RA) a risk factor for severe COVID-19? To answer this question, researchers are unraveling the molecular mechanisms that link the two diseases.

💾 Public Data: Public gene expression dataset (GEO)
Similar to the previous featured study, this one combined data from two public gene expression datasets - one on COVID-19, and another on RA. The researchers found that there were certain genes commonly expressed in patients with both diseases (103 in total). This research team analyzed the data and identified a set of ‘hub genes’ implicated in the biology of both diseases. They also identified a set of candidate drug compounds, already used to treat other diseases, that could be effective at targeting the proteins encoded by the hub genes.
Developing approaches to target the genes driving severe COVID symptoms in people with RA could save lives. Further research at the bench can investigate each ‘hub gene’ and its specific role in disease more closely - and drugs may be designed or repurposed in an attempt to target them.
🗺 Translational Path
Similar Article: Determine the common biological response underlying SARS-CoV-2 infection, IPF, and COPD [Mahmud et al., 2021] 🇨🇳🇧🇩🇦🇺
Cited By: Identify immune system biomarkers for diabetic cardiomyopathy (DCM) based on public data, verify that these markers are elevated in DCM with animal models, and use computer simulations to test known drug compounds against these markers [Guo et al., 2022] 🇨🇳
Cited By: Use computational methods to screen a series of terpenoids (medicinal plant compounds) for their ability to target human proteins that enable SARS-CoV-2 to enter host cells and the proteins of the virus itself [Gyebi et al., 2021] 🇳🇬🇪🇬
(C) What does COVID-19 do to the gut?
🏘Reader Keywords: #COVID-19 #GI Disease #Transcriptomics #Hub Genes #Hepatitis B #Liver Cancer #Nicotinamide
Featured Article: Identification of Key Pathways and Genes in SARS-CoV-2 Infecting Human Intestines by Bioinformatics Analysis 🇨🇳
COVID-19 is not just a lung disease - many infected people have GI symptoms as well. What explains the gut-specific effects of coronavirus infection?

💾 Public Data: Public gene expression dataset (GEO)
Much like the other studies featured in this edition, researchers examined public data on intestinal organoids - small lab-grown structures that resemble intestinal tissue - exposed to coronavirus at different time points. Again, the researchers identified a set of hub genes that seem to govern the response to infection.
What is to be done with these hub genes? Researchers can build on the findings of the study to determine how the protein products of hub genes may interact with viral proteins - and once this is known, try to develop therapeutics that target these proteins.
🗺 Translational Path
Similar Article: Identify the hub genes associated with hepatitis B virus-associated liver cancer - and then correlate expression of hub genes with patient prognosis [Xie et al., 2021] 🇨🇳
Cited By: Show that the human protein DDX17 interacts with viral protein HBx to promote hepatitis B virus-associated liver cancer [Dong et al., 2022] 🇨🇳
Similar Article: Show that the human protein SIRT1 also interacts with viral protein HBx to promote hepatitis B virus-associated liver cancer. Further show that the known SIRT1 inhibitor Nicotinamide can suppress viral replication [Wang et al., 2020] 🇨🇳
(D) The Gut Microbiome Evolves with the Stages of Liver Disease
🏘Reader Keywords: #Hepatitis B #Liver Cirrhosis #Liver Cancer #Microbiome Analysis #Gut Microbiome #Probiotics #Microbial Translocation #HIV #Inflammation
Featured Article: Gut Microbiome Signatures in the Progression of Hepatitis B Virus-Induced Liver Disease [Li et al., 2022] 🇨🇳
Hepatitis B virus (HBV) infection can lead to liver cancer through multiple stages of disease (chronic hepatitis B → liver cirrhosis → cancer). Researchers analyzed how the gut microbiome changes along the way - and whether it might play a role in disease progression.

The research team analyzed nearly 500 public fecal microbiome samples and nearly 100 samples from their own lab representing all stages of liver disease (from chronic hepatitis B to cancer). Looking at all the samples together, they found consistent patterns: certain microbes are elevated at each stage of disease (for example - Lachnospiraceae bacteria in people with chronic hepatitis B), and others are depleted (For example - Prevotella and Oscillibacter bacteria in liver cirrhosis; Coprococcus and Faecalibacterium in liver cancer).
💾 Public Data: Public 16S sequencing (microbiome) datasets (SRA)
This information about how the microbiome changes through the progression to HBV-induced liver cancer offers a new way to monitor disease progression, and to treat patients more effectively. Therapies may even be designed that modify the microbiome as a way to slow or stop cancer progression.
🗺 Translational Path:
Similar Article: Show how the microbiome changes in the progression from a healthy state to initial infection with HBV, through chronic hepatitis B to liver cirrhosis. [Chen et al., 2020] 🇨🇳🇺🇸
Cited By: Treat patients with probiotics to reduce the movement of gut bacteria to other body sites (‘microbial translocation’), a complication of cirrhosis. Gut microbiome changes in cirrhosis patients (such as those noted in the featured study and the similar article above) are known to contribute to bacterial translocation. [Maslennikov et al., 2022] 🇷🇺
Similar Article: Show how probiotic treatment for HIV patients changes the gut microbiome and also reduces microbial translocation and symptoms of chronic inflammation. [Villar-Garcia et al., 2017] 🇪🇸
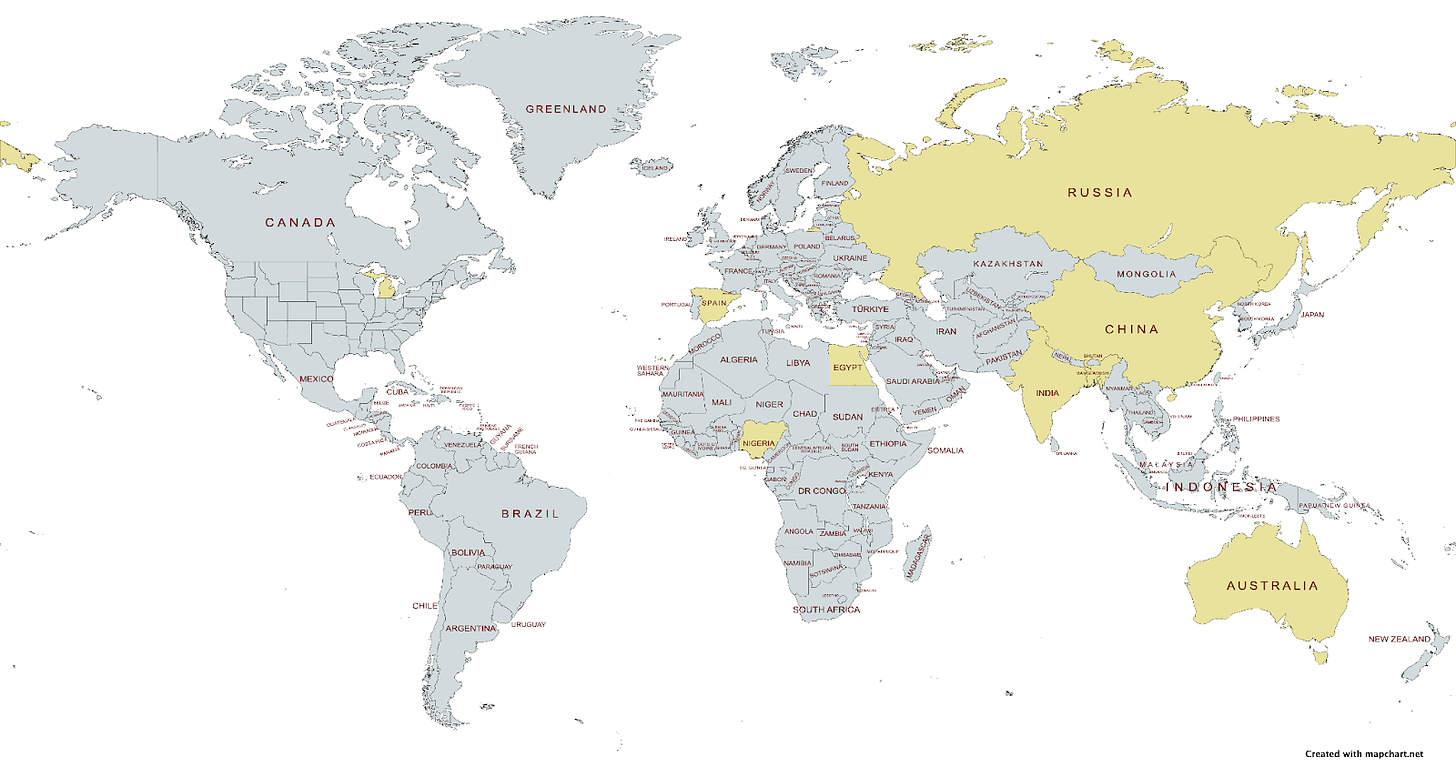
Research Community
This month’s featured research involved 9 countries, including 1 US state!